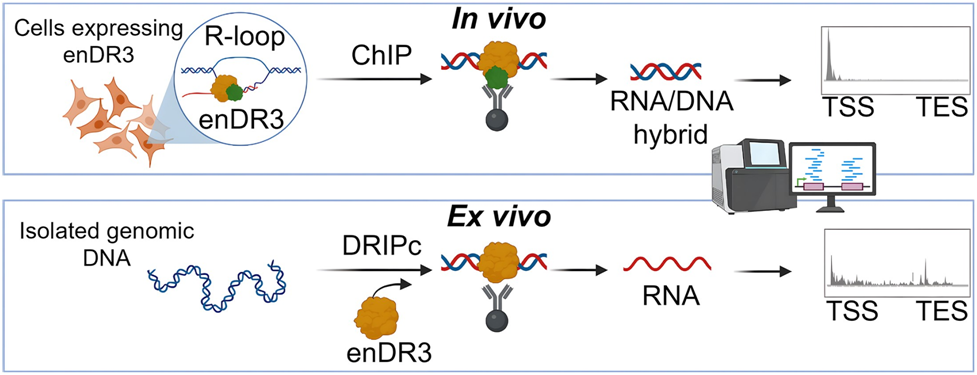
In vivo and ex vivo detection of RNA/DNA hybrids
Researchers from the Institute of Biochemistry and Biophysics PAS and the International Institute of Molecular and Cell Biology have developed a new method for detecting RNA/DNA hybrids found in R-loop structures.
R-loops are crucial nucleic acid structures that occur when a newly transcribed RNA molecule hybridizes with a complementary DNA strand. This process creates an RNA/DNA hybrid and displaces a single-stranded DNA segment. R-loops play a significant role in various biological pathways, and their abnormal formation can lead to genome instability, making them an important topic of study. To investigate R-loops effectively, it is essential to monitor their formation and map their locations within the genome. Until now, many methods for detecting and isolating R-loops have relied on S9.6 monoclonal antibodies, which have proven to be inadequate.
A collaborative effort from two research teams led by Roman Szczęsny (IBB PAS) and Marcin Nowotny (IIMCB) has resulted in the development of a rationally designed RNA/DNA hybrid-binding protein named enDR3 (engineered N-terminal domain of RNase H3). This protein is based on the small N-terminal hybrid-binding domain of the bacterial enzyme RNase H3. The researchers created a tandem version of this domain and introduced a specific mutation to enhance its affinity and specificity for RNA/DNA hybrids. They biochemically characterized this protein and demonstrated its usefulness for specific mapping of RNA/DNA hybrids throughout the human genome, both in ex vivo conditions (isolated nucleic acids) and within a cellular context.
This innovative method has been detailed in a patent application (P.445172) and a recent article published in Nucleic Acids Research.
Genome-wide in vivo and ex vivo mapping of R-loops using engineered N-terminal hybrid-binding domain of RNase H3 (enDR3). Jedynak-Slyvka M, Kaczmarska Z, Graczyk D, Jurkiewicz A, Figiel M, Borowski LS, Szczesny RJ, Nowotny M. Nucleic Acids Res. 2025.